- Biological Products Analysis Service
- Preclinical Drug Development
- Solutions
- High-Throughout Gene Knockout Services
- Biopharmaceutical Stability Analysis Service
- PROTAC Drug Off-Target Protein Assessment Service
- Biosimilar Assessment Service
- Generic Peptide Pharmaceuticals Assessment Service
- Research on Media Development and Optimization
- Small Molecule Drug Target Identification and Validation Service
- Vaccine Quality Research and Testing Solutions
- Antibody Drug Analysis Service
- mRNA Drug Analysis Service
- Medical Device Analysis Service
- Proteomics Analysis Service
- Sample Preparation Service
- Protein Analysis Service
- Quantitative Proteomics Service
- Post-Translational Modifications Proteomics Service
- Proteomics Service of Different Sample Types
- Targeted Proteomics Service
- Peptidomics Analysis Service
- Proteomics Bioinformatic Analysis Service
- High-Throughout Proteomics Analysis Service for Gene Knockout
- Olink Analysis Service
- Chemical Proteomics Analysis Service
- Metabolomics Analysis Service
- Lipidomics Analysis Service
- Multi-Omics Analysis Service
- Integrative Lipidomics-Proteomics Analysis Service
- Integrative Transcriptomics-Metabolomics Analysis Service
- Integrative Transcriptomics-Proteomics Analysis Service
- Integrative Transcriptomics-Lipidomics Analysis Service
- Integrative Metabolomics-16S rDNA Sequencing Analysis Service
- Integrative Proteomics-Metabolomics Analysis Service
- Integrative PTM-Metabolomics Analysis
- Transcriptome Sequencing (RNA-sequencing) Service
- Metatranscriptomics Sequencing Service
- Full-Length 16S/18S/ITS Sequencing Service
- Prokaryotic Transcriptomics Sequencing Service
- Eukaryotic Non-Ginseng Transcriptome Sequencing Service
- Eukaryotic Transcriptome Sequencing Service
- 16S/18S/ITS Amplicon Sequencing Service
- Exosomal Sequencing Service
- Single Cell Analysis Service
- Bioinformatics
- Proteomics Analysis Service
SILAC Based Co-IP-MS for Protein Interaction Analysis Service
Co-immunoprecipitation (Co-IP) is a classic method based on the principle that antibodies specifically recognize particular antigens for studying protein-protein interactions. Co-IP is commonly used to examine protein interactions under physiological conditions, and when combined with mass spectrometry (MS), it can be used to identify novel interacting proteins. MtoZ Biolabs conducted Co-IP experiments by labeling experimental and control cell groups using the SILAC method. Immunocomplexes were isolated based on the specific reaction between antigens and antibodies. Qualitative and quantitative detection of proteins in the immunocomplexes was performed using LC-MS/MS. When the content of a protein in the experimental group differs statistically from that in the control group, it is determined that this protein interacts with the protein under study, greatly reducing the possibility of false-positive results in protein interaction. The combination of SILAC and Co-IP-MS techniques can be used for high-throughput quantitative analysis of protein interaction networks under specific conditions.
The SILAC approach introduces mass differences from the source, reducing experimental system errors and providing rich quantitative information with high accuracy. MtoZ Biolabs utilizes Thermo Fisher's Q Exactive HF, Orbitrap Fusion, and Orbitrap Fusion Lumos mass spectrometry platforms combined with Nano-LC to introduce the SILAC-based Co-IP-MS service package. Simply inform us of your experimental objectives and send us your cells, and we will take care of all subsequent project-related matters, including cell culture, cell labeling, protein extraction, antibody IP, efficiency detection, protein digestion, peptide separation, mass spectrometry analysis, raw data analysis, and bioinformatics analysis.
Analysis Workflow
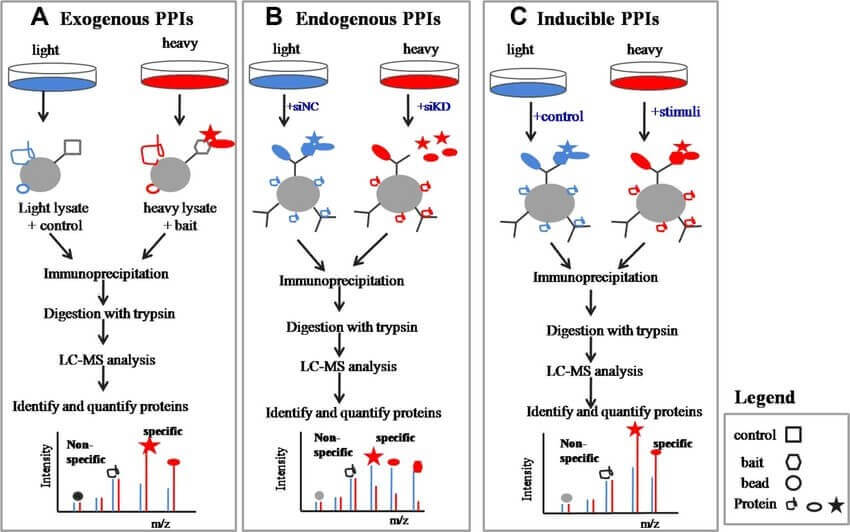
Chen, X. L. et al. Proteomics. 2015.
Figure 1. SILAC-Based Co-IP-MS Experimental Workflow
Service Advantages
1. Preservation of Protein Structure and Post-Translational Modifications as much as Possible
2. Protein Interactions Occur in Their Natural States, Avoiding Artificial Influences
3. Natural Interaction Protein Complexes can be Isolated
4. High-throughput Analysis Enables Rapid and Comprehensive Analysis of All Interacting Proteins
5. Contrast with Control Groups to Identify Non-Specific Binding Proteins, Minimizing False Positives
6. Comparative Analysis can Determine the Relative Abundance of Each Interacting Protein
Sample Submission Requirements
1. Simply Send us Frozen Cells Using Dry Ice, Ensuring that the Cells can be Passaged at least 5 Times
2. If you Need to Label Cells Yourself and Complete Processes such as Cell Stimulation, Simply Send us the Labeled and Processed Cell or Protein Samples on Dry Ice. For Each Sample, Ensure a Total Protein Content of at least 300 μg
Note: Please transport samples using an adequate amount of dry ice and choose a faster shipping method to minimize the possibility of sample degradation during transport.
Deliverables
In the technical report, MtoZ Biolabs will provide you with detailed technical information, including:
1. Experimental Procedures
2. Relevant Mass Spectrometry Parameters
3. Detailed Information on Identified Proteins
4. Mass Spectrometry Images
5. Raw Data
6. Bioinformatics Analysis
How to order?



